PACER: Acyclic Causal Discovery from Large-scale Interventional Data
PACER (Perturbation-driven Acyclic Causal Edge Recovery) is a scalable framework for causal discovery from interventional data that guarantees acyclicity by design.
Modern technologies like single-cell Perturb-seq now generate interventional data across thousands of genes, but existing differentiable discovery methods exhibit limited performance and scalability. These methods typically treat acyclicity as a soft penalty rather than a hard constraint, leading to numerical instability, high computational costs, and optimization over cyclic configurations that lack a well-defined joint likelihood. PACER addresses this by redesigning the search space so that every explored graph is guaranteed to be a valid, acyclic structure by design, avoiding expensive regularization terms and enabling faster and more accurate causal discovery in large-scale datasets.
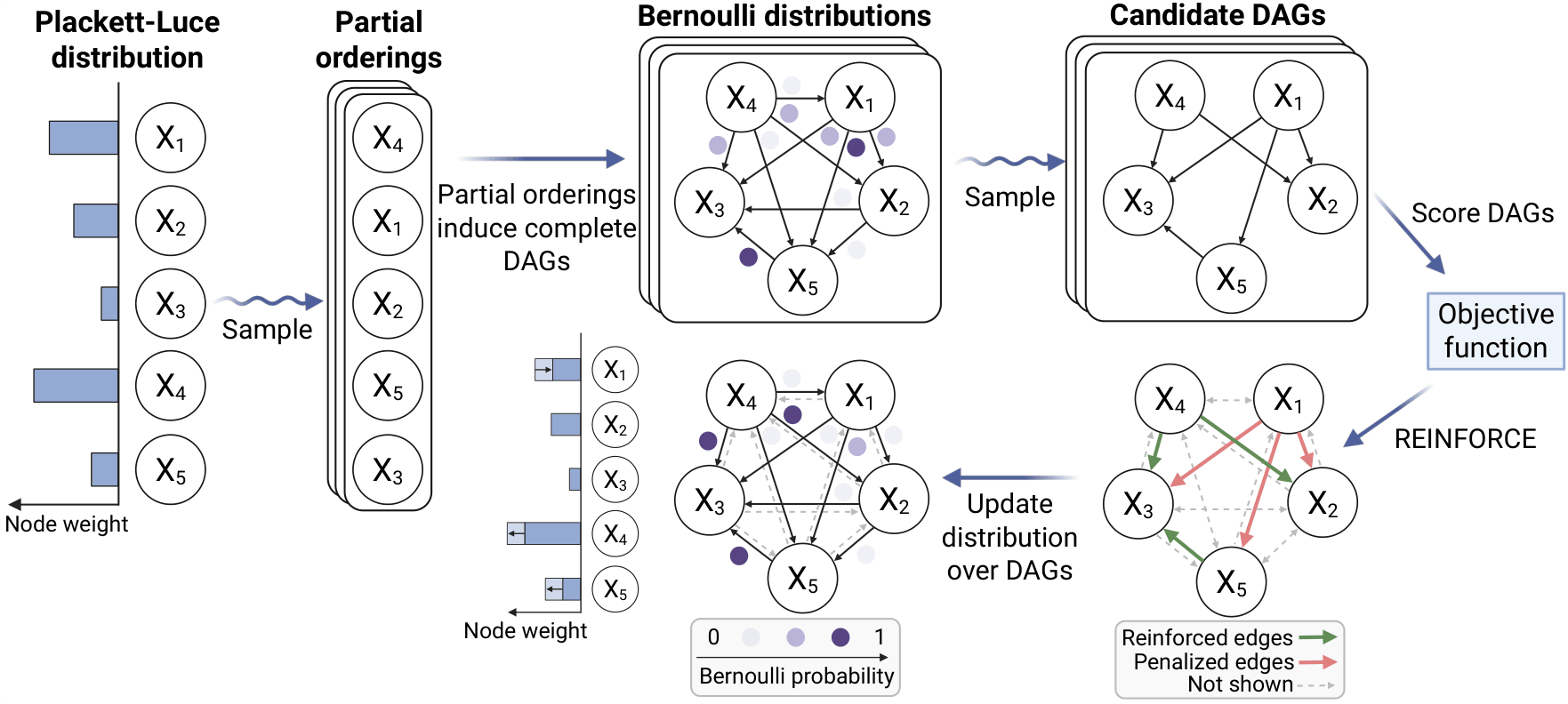
PACER models a topological ordering of variables using a Plackett-Luce distribution. Nodes with higher weight are more likely to precede nodes with lower weight in downstream DAGs. Samples from this distribution induce complete DAGs, which are further filtered via samples from independent, edge-specific Bernoulli distributions. This defines our Bernoulli-Plackett-Luce distribution over DAGs. At train time, we sample multiple candidate graphs and score them based on a likelihood-based objective function. We then optimize the parameters of the Bernoulli-Plackett-Luce model using either an analytic estimator or REINFORCE gradient updates.
The key features of PACER are:
- Acyclicity by design. We introduce a distribution over DAGs to simultaneously model topological orderings and filter causal edges. This formulation ensures that every structure evaluated during optimization is strictly acyclic, eliminating the need for expensive acyclicity terms and ensuring that the search remains within the space of DAGs throughout the entire learning process.
- Flexible differentiable causal discovery. PACER provides a unified likelihood-based framework that jointly leverages observational and interventional data. This approach supports a variety of conditional density models, including neural parameterizations and tailored distributions for specific data modalities.
- Exact gradient computation. We derive a closed-form expression for an interventional log-likelihood objective under linear-Gaussian mechanisms. This analytic gradient enables PACER to scale to thousands of variables, achieving up to a two-order-of-magnitude speedup over penalty-based differentiable methods.
- Inductive biases and prior knowledge. PACER is a flexible framework that supports the direct integration of structural priors, such as node centrality expectations or transcription factor binding constraints when modeling genomic regulatory networks.
Empirically, PACER matches or exceeds the performance of state-of-the-art causal discovery methods on protein signaling and large-scale genetic perturbation benchmarks, while scaling efficiently to networks with thousands of variables. In particular, the analytic formulation for linear-Gaussian mechanisms yields substantial computational gains, demonstrating that exact and scalable causal discovery is achievable when the search space is appropriately structured. Taken together, these results position PACER as a practical solution for causal discovery from interventional data in modern high-dimensional settings that remain challenging for existing methods.
Publication
PACER: Acyclic Causal Discovery from Large-scale Interventional Data
Ramon Viñas Torné, Sílvia Fábregas Salazar, Soyon Park, Ivo Alexander Ban, Artyom Gadetsky, Nikita Doikov, Maria Brbić.
@article{vinas2026pacer,
title = {PACER: Acyclic Causal Discovery from Large-scale Interventional Data},
author = {Vi{\~n}as Torn{\'e}, Ramon and F{\`a}bregas Salazar, S\'{\i}lvia and Park, Soyon and Ban, Ivo Alexander and Gadetsky, Artyom and Doikov, Nikita and Brbi{\'c}, Maria},
journal = {Proceedings of the 43rd International Conference on Machine Learning (ICML)},
year = {2026},
}
Code
An implementation is available on GitHub.
Contributors
The following people contributed to this work:
Ramon Viñas Torné
Sílvia Fábregas Salazar
Soyon Park
Ivo Alexander Ban
Artyom Gadetsky
Nikita Doikov